Spatial Transcriptomics with Automated Fluidic Control
Plug-and-play fluidic automation for merfish/seqfish/seqfish+
Instruments for spatial transcriptomics
The perfect pack for your fluorescence in situ hybridization setup
Intergration and automation
Perform automated personalized sequences and microscope synchronisation through TTL triggers and SDK
Easy sequential injection
Quickly swap between solutions with only one flow controller

Need a microfluidic SME partner for your Horizon Europe project?
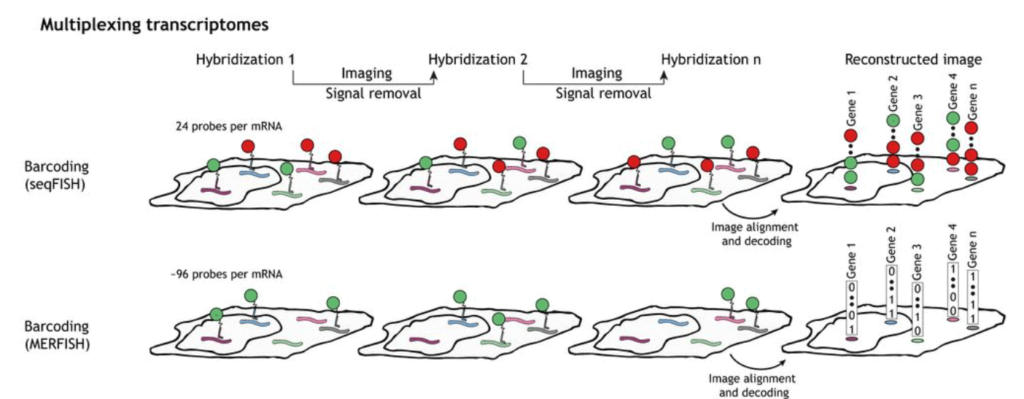
Sequential hybridization figures by Lubeck et al. (2014) and Shah et al. (2016)
Fluidic spatial transcriptomics
Quickly implement your MERFISH/seqFISH/seqFISH+ experiments with this all-included, user-friendly, customizable, and automatable spatial transcriptomics pack.
A state-of-the-art flow controller, coupled with other instruments, like selecting valves, is ideally suited to precisely and automatically dispense tens of dyes, imaging, and washing solutions. This pack is more cost-efficient, practical, and easy-to-use than syringe pumps. This platform is also highly flexible and versatile, as it can quickly adapt to the FISH experiment’s required solutions.
This pack aims to allow you to perform multiplexed fluorescence in situ hybridization at a microfluidic scale, significantly reducing the cost of each experiment by reducing the amount of solution. It also allows the system to be automated and improves the reproducibility of results.
This easy-to-use microfluidic spatial transcriptomics for MERFISH/seqFISH pack contains several critical functions, including a precise and perfectly controlled flow rate for precise small dispensed volume, a fast and easy sequential injection between different solutions, and flexible software for sequence setting and automation with the built-in automation sequencer.
The sequence scheduler transforms the setup into an all-in-one automatized platform that can easily flow many different solutions for your spatial transcriptomics and can further be synchronized with other scientific equipment, such as a fluorescence microscope.
Contact our experts for help integrating your experiment using TTL triggers or direct software integration via SDK.
A microfluidic chip is used to perform spatial transcriptomics. Chips are transparent, available in several materials and shapes, with different numbers of inlets/outlets, and compatible with varying FISH methods. We can provide different chips with this pack with specific channel heights, widths, lengths, and materials. Spatial transcriptomics analysis can be expanded even further by using several microfluidic chips in parallel or one microfluidic chip with several channels.
Our all-in-one pack guarantees good compatibility between the different instruments, allows you to start your experiment right away, is piloted by a single software, and can be used for other applications. We also provide continuous and full customer support to fulfill your experiment goals.
Spatial transcriptomics pack setup
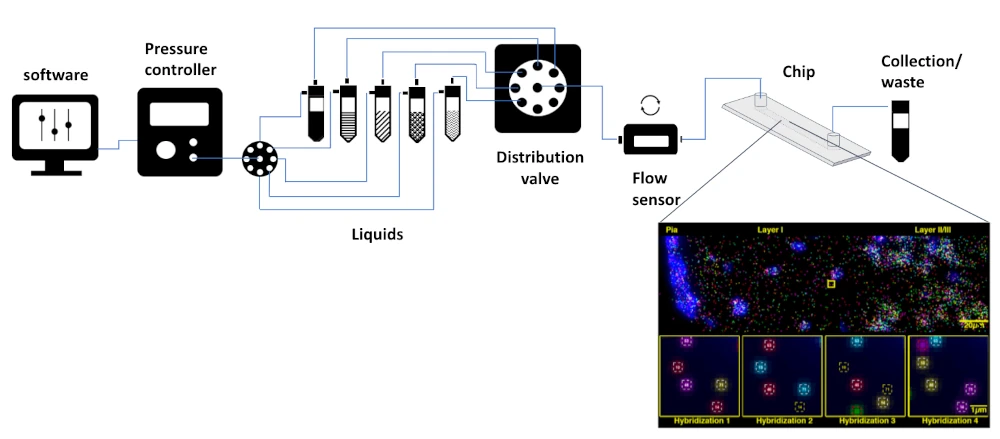
A typical pack contains:
Flow sensor (Galileo, MIC)
Software (Galileo user interface)
Pressure controller
Distribution valve
Tubings and luers
Several Eppendorfs or Falcon reservoirs
Microfluidic chips
A user guide
Spatial transcriptomics principle
The field of research that studies spatially resolved transcriptomes has recently been gaining interest to expand our knowledge of complex multicellular biological systems and their interactions [2]. It has also been chosen by Nature Methods as the “Method of the Year” for 2020 [3].
Cells can sense neighboring cells and their local environment through molecular and cellular mechanisms; thus, it is crucial to study complex biological phenomena to measure the phenotypic and genomic states of single cells in their respective spatial positions [4].
SeqFISH and MERFISH are in situ hybridization encoding methods that use probe detection for single-cell spatial transcriptomics [5]. They are an improved version of the smFISH technique used to study gene expression variations and their consequences [6]. Other methods have also been developed at an increasing rate in the last few years to obtain spatially resolved transcriptomes [7].
To perform seqFISH, a first in situ hybridization is done with one set of fluorescent FISH probes and a labeled dye. DNase is then used to remove the fluorophores. The mRNA is hybridized with the identical FISH probes but labeled with another dye. With several dyes and rounds of hybridization, it is possible to barcode many genes in single cells [8].
SeqFISH has been improved to seqFISH+, which can be used for cell spatial and biological process studies by combining seqFISH with a confocal microscope. This was used to perform super-resolution imaging and multiplexing of 10,000 genes in a single cell [9].
Multiplexed Error Robust Fluorescence In Situ Hybridization (MERFISH) is an improvement to the single-molecule Fluorescence In Situ Hybridization (smFISH) method by massively parallelizing and identifying hundreds to thousands of RNA species simultaneously with spatial information.
This method is also error-robust, thanks to using unassigned binary barcodes that can detect and correct errors. This is the main difference between it and seqFISH, coded in a color sequence [10].
Using lab-on-chip technologies and microfluidic platforms provides several improvements to seqFISH and MERFISH. Mainly to reduce the cost, which can be pretty high for these methods, the experiment time, and to automate the process [11].
References
Shah, Sheel & Lubeck, Eric & Zhou, Wen & Cai, Long. (2016). In Situ Transcription Profiling of Single Cells Reveals Spatial Organization of Cells in the Mouse Hippocampus. Neuron. 92. 342-357.
Asp M., Bergenstrahle J., Lundeberg J., Spatially Resolved Transcriptomes—Next Generation Tools for Tissue Exploration, BioEssays, 2020, 42 (10)
Marx, V. Method of the Year: spatially resolved transcriptomics. Nat Methods 18, 9–14 (2021).
Mayr U., Serra D., Liberali P. Exploring single cells in space and time during tissue development, homeostasis and regeneration. Development, 2019, 146(12),
Zhou Y, Jia E, Pan M, Zhao X, Ge Q. Encoding Method of Single-cell Spatial Transcriptomics Sequencing. Int J Biol Sci 2020; 16(14):2663-2674.
Raj A, van Oudenaarden A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell. 2008;135:216–226.
Asp, M., Bergenstråhle, J., Lundeberg, J., Spatially Resolved Transcriptomes—Next Generation Tools for Tissue Exploration. BioEssays 2020, 42, 1900221.
Lubeck, E., Coskun, A., Zhiyentayev, T. et al. Single-cell in situ RNA profiling by sequential hybridization. Nat Methods 11, 360–361 (2014).
Eng, CH.L., Lawson, M., Zhu, Q. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 568, 235–239 (2019).
Moffitt, J R, and X Zhuang. “RNA Imaging with Multiplexed Error-Robust Fluorescence In Situ Hybridization (MERFISH).” Methods in enzymology vol. 572 (2016): 1-49.
Rodriguez-Mateos, P., Azevedo, N.F., Almeida, C. et al. FISH and chips: a review of microfluidic platforms for FISH analysis. Med Microbiol Immunol 209, 373–391 (2020).
Why use microfluidics for fluorecence in situ hybridization?
Using microfluidics is the most efficient method to perform MERFISH (Multiplexed Error-Robust Fluorescence In Situ Hybridization) or seqFISH (sequential Fluorescence In Situ Hybridization) and observe multiple genes and their spatial configuration. It permits experimentation with a drastically lower amount of expensive dye and buffer solutions and is compatible with biological applications and microscope observations.
As mentioned above, an automatized sequence can be implemented to inject the solutions into the cell. Furthermore, several different chips can be connected to the system to observe different samples in parallel easily.
This pack can also be combined with other microfluidic steps before this fluorescent in situ hybridization setup; for single-cell isolation, for example, you can use microfluidic single-cell encapsulation [1].
Microfluidics can also be used for the MA-FISH method, which uses oscillatory flows of diluted probe solutions or performs barcoding (DBiT-seq).
The Microfluidics Innovation Center has developed expertise in microfluidics over more than ten years. It can provide its state-of-the-art expertise in biology and engineering, making it the perfect partner for your transition to microfluidics.
References
1. Mayr U., Serra D., Liberali P. Exploring single cells in space and time during tissue development, homeostasis and regeneration. Development, 2019, 146(12),
Customize your fluidic control for spatial transcriptomics pack
This pack is highly customizable to fit your experiment specification and spatial transcriptomics methods (MERFISH, seqFISH,…).
If you’re unsure about the instrument choices and settings best suited for your application, contact one of our experts!

Funding and Support
The ElectroMed project results helped develop this pack, funded by the European Union’s Horizon 2020 FETOPEN under grant agreement No 862539.
Products & Associated Accessories
FAQ - Spatial transcriptomics with automated fluidic control
What is the Fluidic Control for Spatial Transcriptomics pack?
The pack is an all-in-one, plug-and-play microfluidic platform designed to automate fluid delivery during fluorescence in situ hybridization (FISH) experiments such as MERFISH and seqFISH/seqFISH+. It replaces manual pipetting and bulky syringe pumps with a precision flow controller, distribution valve, and integrated software, enabling researchers to sequentially inject dyes, imaging buffers, and washing solutions into a microfluidic chip in a controlled, reproducible manner. The system is piloted by a single interface and can be synchronized with fluorescence microscopes via TTL triggers or SDK integration.
What spatial transcriptomics techniques can be used with this pack?
The pack is mostly targeted at:
-MERFISH (Multiplexed Error-Robust FISH)- allows individual simultaneous detection of hundreds to thousands of RNA species with localization information,
-seqFISH (sequential FISH) – sequentially hybridizes genes with various rounds of different fluorescent dyes,
-seqFISH+ is a super-resolution imaging method using confocal microscopy that improves upon it by sequencing up to 10,000 genes per cell.
The pack can also be used with MA-FISH (oscillatory flow-based probe delivery and DBiT-seq barcoding workflows.
What are the instruments that are added in a standard pack setup?
-A pressure-driven flow controller to provide precise and pulse-free flow,
-A real-time flow sensor (Galileo, MIC),
-A distribution valve for fast sequential switching between solutions,
-Tubing and luer, which are used to assemble the system,
-Eppendorf or Falcon tube reservoirs,
-Microfluidic chips adapted to your FISH method,
-The Galileo user interface software.
-A user guide for setup and operation.
Components are all pre-tested to ensure compatibility, and experiments can start as soon as the components arrive.
Why is the use of pressure-based flow control desirable compared to the use of syringe pumps in FISH experiments?
The pressure controllers have a number of benefits over the traditional syringe pumps in the FISH workflow:
-Quicker conversion of solutions with a small dead volume between injections,
-Laminar flow without pulse and keeping fragile biological samples intact and having equal distribution of the probe,
-Increased cost-efficiency, the system will use less expensive fluorescent dyes and buffers per experiment.
-Easier to use, fewer moving parts, and integrated software interface,
This improves scalability, allowing multiple chips to be connected in parallel to run samples in parallel.
What is the way automation operates in this system and what can it control?
It has an in-built sequence scheduler (Galileo user interface) that eases programmability with fully automated injection sequences. The researchers can establish the sequence, timing, and flow rate for each solution, and then perform the entire process remotely. Major automation characteristics are:
-Multi-round hybridization with customizable injection sequences,
-Synchronization with fluorescence microscopes via TTL triggers for timed image acquisition,
-Direct SDK integration for software-level control from third parties.
-Parallel chip management of experimental high-throughput design.
This converts the apparatus into a completely automated and reproducible system for long, multi-cycle FISH procedures.
What are the types of microfluidic chips, and how are they chosen?
Chips are clear and can be made of various materials (e.g., PDMS, glass, polycarbonate) and geometries and have channel heights, widths, and lengths that can be customized. Selection depends on:
-The FISH technique employed (seqFISH, MERFISH etc.),
-The type of biological sample (tissue section, dissociated cells, etc.),
-The required inlets and outlets to be used in sequential injections,
-Conformability with the microscope objective and modality.
There are several ways to operate multiple chips in parallel, either using a single flow controller or a single chip with many channels to handle multiple samples. MIC professionals can provide the advice on the best chip configuration to use in your experiment.
What is the cost reduction of spatial transcriptomics experiments with this pack?
The cost of MERFISH and seqFISH workflows is high because many fluorescently labeled probes and buffers are used across several rounds of hybridizations. This pack handles the issue of cost in the following ways:
-Shifts reactions to nanoliter-to-microliter scale microchannels and used by reducing to a minimum the amount of the reagent required per experiment,
-Eliminates the error inherent in manual pipetting, resulting in expensive loss of samples or failed experiment,
-Enhances reproducibility so that experiments do not have to be repeated again,
-Allows multiplexing between parallel chips, to put maximum output on each reagent batch.
This renders the system especially appealing to laboratories conducting spatial transcriptomics investigations, which are high-frequency or high-gene-count.
Is the pack compatible with established laboratory systems and microscopes?
Yes. The pack can be interfaced with the existing fluorescence microscopy infrastructure. The choices of integration comprise:
-Hardware-level synchronization TTL triggers with acquisition of images on a microscope,
-Custom software pipeline SDK access, instrument control SDK access,
-Tubing connections that can fit standard lab fittings,
-The compatibility of upstream microfluidics steps, including single-cell encapsulation/isolation modules.
The MIC engineering team fully supports custom integration, including recommendations on how to get the system to work with inverted microscopes, confocal systems, or home-built imaging systems.
How would the Microfluidics Innovation Center be useful as a consortium partner in Horizon Europe?
MIC is a skilled SME with more than a decade of experience in microfluidics engineering and biological applications, making it a good partner for Horizon Europe research projects involving lab-on-chip technologies. MIC can contribute to a consortium in the following ways:
-Non-academic beneficiary or associated partner offering special fluidic instrumentation and integration skills,
-A work package work technology provider specializing in automated fluid handling, chip design, or FISH-based assay development,
-A joint developer of automated microfluidic application systems was tested in the project’s pipeline.
How customizable is the pack, and what support does MIC give?
It is highly modular and can be adapted to any spatial transcriptomics workflow. The customization options are:
-Setting flow rate ranges and pressure settings to particular probe viscosities or chip geometries,
-Choosing chips with channel dimensions and materials that are custom,
-Increasing the number of solution reservoirs in the valve construction,
-Compiling the software automation sequences to the timing and trigger requirements of the experiments.
MIC will accompany you throughout the installation process, up to the actual execution of the experiment, including troubleshooting, protocol optimization of their systems, and expert consultation.

